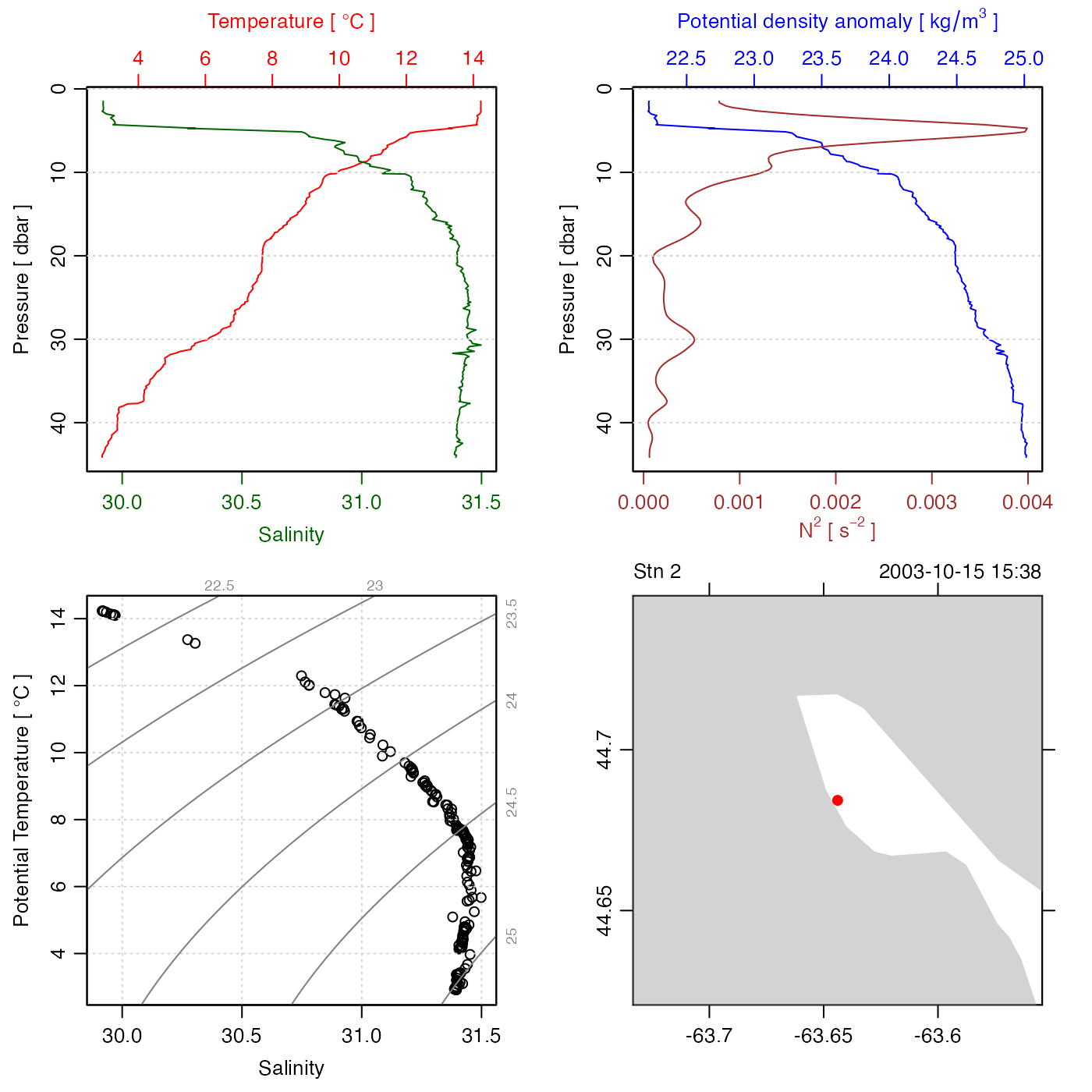
Plot CTD data in any of many different ways. In many cases, the best choice is to use default values for all parameters other than the first. This yields a 4-panel plot that displays a basic overview of the data, with a combined profile of salinity and temperature at the top left, a combined plot of density and the square of buoyancy frequency at top right, a TS diagram at bottom left, and a map at bottom right.
Usage
# S4 method for class 'ctd'
plot(
x,
which,
col = par("fg"),
fill,
borderCoastline = NA,
colCoastline = "lightgray",
eos = getOption("oceEOS", default = "gsw"),
ref.lat = NaN,
ref.lon = NaN,
grid = TRUE,
coastline = "best",
Slim,
Clim,
Tlim,
plim,
densitylim,
sigmalim,
N2lim,
Rrholim,
dpdtlim,
timelim,
drawIsobaths = FALSE,
clongitude,
clatitude,
span,
showHemi = TRUE,
lonlabels = TRUE,
latlabels = TRUE,
latlon.pch = 20,
latlon.cex = 1.5,
latlon.col = "red",
projection = NULL,
cex = 1,
cex.axis = par("cex.axis"),
pch = 1,
useSmoothScatter = FALSE,
df,
keepNA = FALSE,
type,
mgp = getOption("oceMgp"),
mar = c(mgp[1] + 1.5, mgp[1] + 1.5, mgp[1] + 1.5, mgp[1] + 1),
inset = FALSE,
add = FALSE,
debug = getOption("oceDebug"),
...
)Arguments
- x
a ctd object.
- which
a numeric or character vector specifying desired plot types. If
whichis not supplied, a default will be used. This default depends ondeploymentTypein themetadataslot ofx. IfdeploymentTypeis"profile"or missing, thenwhichdefaults toc(1, 2, 3, 5). IfdeploymentTypeis"moored"or"thermosalinograph"thenwhichdefaults toc(30, 3, 31, 5). Finally, ifdeploymentTypeistowyothenwhichdefaults toc(30, 31, 32, 3).The details of individual
whichvalues are as follows. Some of the entries refer to the EOS (equation of state for seawater), which may either"gsw"for the modern Gibbs Seawater system, or"unesco"for the older UNESCO system. The EOS may be set with theeosargument toplot,ctd-method()or by usingoptions(), withoptions(oceEOS="unesco")oroptions(oceEOS="unesco"). The default EOS is"gsw".which=1orwhich="salinity+temperature"gives a combined profile of temperature and salinity. If the EOS is"gsw"then Conservative Temperature and Absolute Salinity are shown; otherwise in-situ temperature and practical salinity are shown.which=2orwhich="density+N2"gives a combined profile of density anomaly, computed withswSigma0(), along with the square of the buoyancy frequency, computed withswN2(). Theeosparameter is passed to each of these functions, so the desired EOS is used.which=3orwhich="TS"gives a TS plot. If the EOS is"gsw", T is Conservative Temperature and S is Absolute Salinity; otherwise, they are in-situ temperature and practical salinity, respectively.which=4orwhich="text"gives a textual summary of some aspects of the data.which=5orwhich="map"gives a map plotted withplot,coastline-method(), with a dot for the station location. Notes near the top boundary of the map give the station number, the sampling date, and the name of the chief scientist, if these are known. Note that the longitude will be converted to a value between -180 and 180 before plotting. (See also notes aboutspan.)which=5.1as forwhich=5, except that the file name is drawn above the map.which=6orwhich="density+dpdt"gives a profile of density and \(dP/dt\), which is useful for evaluating whether the instrument is dropping properly through the water column. If the EOS is"gsw"then \(\sigma_0\) is shown; otherwise, \(\sigma_\theta\) is shown.which=7orwhich="density+time"gives a profile of density and time.which=8orwhich="index"gives a profile of index number, which can provide useful information for trimming withctdTrim().which=9orwhich="salinity"gives a profile of Absolute Salinity if the EOS is"gsw", or practical salinity otherwise.which=10orwhich="temperature"gives a profile of Conservative Temperature if the EOS is"gsw", or in-situ temperature otherwise.which=11orwhich="density"gives a profile of density as computed withswRho(), to which theeosparameter is passed.which=12orwhich="N2"gives an \(N^2\) profile.which=13orwhich="spice"gives a profile of the UNESCO-defined spice variable.which=14orwhich="tritium"gives a tritium profile.which=15orwhich="Rrho"gives a diffusive-case density ratio profile.which=16orwhich="RrhoSF"gives a salt-finger case density ratio profile.which=17orwhich="conductivity"gives a conductivity profile.which=20orwhich="CT"gives a profile of Conservative Temperature.which=21orwhich="SA"gives a profile of Absolute Salinity.which=30orwhich="Sts"gives a time series of Salinity Absolute Salinity if the EOS is"gsw"or practical salinity otherwise.which=31orwhich="Tts"gives a time series of Conservative Temperature if the EOS is"gsw"or in-situ temperature otherwise.which=32orwhich="pts"gives a time series of pressurewhich=33orwhich="rhots"gives a time series of density anomaly, \(\sigma_0\) if the EOS is"gsw"or \(\sigma_\theta\) otherwise.otherwise,
whichis interpreted as a character value to be checked against thedataanddataDerivedfields returned byx[["?"]. If a match is found then a profile of the corresponding quantity is plotted. If there is no match, an error is reported.
- col
color of lines or symbols.
- fill
a legacy parameter that will be permitted only temporarily; see “History”.
- borderCoastline
color of coastlines and international borders, passed to
plot,coastline-method()if a map is included inwhich.- colCoastline
fill color of coastlines and international borders, passed to
plot,coastline-method()if a map is included inwhich. Set toNULLto avoid filling.- eos
character value indicating the equation of state to be used, either
"unesco"or"gsw". The default is to use a value stored withoptions()as e.g.options(oceEOS="unesco").- ref.lat
latitude of reference point for distance calculation. The permitted range is -90 to 90.
- ref.lon
longitude of reference point for distance calculation. The permitted range is -180 to 180.
- grid
logical value indicating whether to draw a grid on the plot.
- coastline
a specification of the coastline to be used for
which="map". This may be a coastline object, whether built-in or supplied by the user, or a character string. If the later, it may be the name of a built-in coastline ("coastlineWorld","coastlineWorldFine", or"coastlineWorldCoarse"), or"best", to choose a suitable coastline for the locale, or"none"to prevent the drawing of a coastline. There is a speed penalty for providingcoastlineas a character string, because it forcesplot,coastline-method()to load it on every call. So, ifplot,coastline-method()is to be called several times for a given coastline, it makes sense to load it in before the first call, and to supply the object as an argument, as opposed to the name of the object.- Slim, Clim, Tlim, plim, densitylim, sigmalim, N2lim, Rrholim, dpdtlim, timelim
optional numeric vectors of length 2, that give axis limits for salinity (or Absolute Salinity, if
eosis"gsw"), conductivity, in-situ or potential temperature (or Conservative Temperature, ifeosis `"gsw"'), pressure, density, density anomaly (either sigma-theta or sigma0), square of buoyancy frequency, density ratio, dp/dt, and time, respectively.- drawIsobaths
logical value indicating whether to draw depth contours on maps, in addition to the coastline. The argument has no effect except for panels in which the value of
whichequals"map"or the equivalent numerical code,5. IfdrawIsobathsisFALSE, then no contours are drawn. IfdrawIsobathsisTRUE, then contours are selected automatically, using pretty(c(0, 300))if the station depth is under 100m or pretty(c(0, 5500))otherwise. IfdrawIsobathsis a numerical vector, then the indicated depths are drawn. For plots drawn withprojectionset toNULL, the contours are added withcontour()and otherwisemapContour()is used. To customize the resultant contours, e.g. setting particular line types or colors, users should call these functions directly (see e.g. Example 2).- clongitude, clatitude, span
controls for the map area view, used only if
which="map".clongitudeandclatitudespecify the centre of the view, andspanspecifies the approximate extend of the view, in kilometres. (Ifspanis not given, it is be determined as a small multiple of the distance to the nearest point of land, in an attempt to show the station in familiar geographical context.)- showHemi, lonlabels, latlabels
controls for axis labelling, used only if
which="map".showHemiis logical value indicating whether to show hemisphere in axis tick labels.lonlabelsandlatlabelsare numeric and character values that control the axis labelling.- latlon.pch, latlon.cex, latlon.col
controls for station location, used only if
which="map".latlon.pchsets the symbol code,latlon.cexsets the character expansion factor, andlatlon.colsets the colour.- projection
controls the map projection (if any), and ignored unless
which="map". The possibilities are as follows. (1) Ifprojection=NULL(the default) then no projection will be used; the map will simply show longitude and latitude in a Cartesian frame, scaled to retain shapes at the centre. (2) Ifprojection="automatic"then either a Mercator or Stereographic projection will be used, depending on whether the CTD station is within 70 degrees of the equator or at higher latitudes. (3) Ifprojection` is a string in the format used bymapPlot(), then it is is passed to that function.- cex
size to be used for plot symbols (see
par()).- cex.axis
size factor for axis labels (see
par()).- pch
code for plotting symbol (see
par()).- useSmoothScatter
logical value indicating whether to use
smoothScatter()instead ofplot()to draw the plot.- df
optional numeric argument that is ignored except for plotting buoyancy frequency; in that case, it is passed to
swN2().- keepNA
logical value indicating whether
NAvalues will yield breaks in lines drawn iftypeisb,l, oro. The default value isFALSE. SettingkeepNAtoTRUEcan be helpful when working with multiple profiles strung together into one ctd object, which otherwise would have extraneous lines joining the deepest point in one profile to the shallowest in the next profile.- type
the type of plot to draw, using the same scheme as
plot(). If supplied, this is increased to be the same length aswhich, if necessary, and then supplied to each of the individual plot calls. If it is not supplied, then those plot calls use defaults (e.g. using a line forplotProfile(), using dots forplotTS(), etc).- mgp
three-element numerical vector specifying axis-label geometry, passed to
par(). The default establishes tighter margins than in the usual R setup.- mar
four-element numerical vector specifying margin geometry, passed to
par(). The default establishes tighter margins than in the usual R setup. Note that the value ofmaris ignored for the map panel of multi-panel maps; instead, the present value of par("mar")is used, which in the default call will make the map plot region equal that of the previously-drawn profiles and TS plot.- inset
logical value indicating whether this function is being used as an inset. The effect is to prevent the present function from adjusting margins, which is necessary because margin adjustment is the basis for the method used by
plotInset().- add
logical value indicating whether to add to an existing plot. This only works if
length(which)=1, and it will yield odd results if the value ofwhichdoes not match that in the previous plots.- debug
an integer specifying whether debugging information is to be printed during the processing. This is a general parameter that is used by many
ocefunctions. Generally, settingdebug=0turns off the printing, while higher values suggest that more information be printed. If one function calls another, it usually reduces the value ofdebugfirst, so that a user can often obtain deeper debugging by specifying higherdebugvalues.- ...
optional arguments passed to plotting functions.
Details
The
default values of which and other arguments are chosen to be useful
for quick overviews of data. However, for detailed work it is common
to call the present function with just a single value of which, e.g.
with four calls to get four panels. The advantage of this is that it provides
much more control over the display, and also it permits the addition of extra
display elements (lines, points, margin notes, etc.) to the individual panels.
Note that panels that draw more than one curve (e.g. which="salinity+temperature"
draws temperature and salinity profiles in one graph), the value of par("usr")
is established by the second profile to have been drawn. Some experimentation will
reveal what this profile is, for each permitted which case, although
it seems unlikely that this will help much ... the simple fact is that drawing two
profiles in one graph is useful for a quick overview, but not useful for e.g. interactive
analysis with locator() to flag bad data, etc.
History of Changes
January 2022:
Add ability to profile anything stored in the
dataslot, and anything that can be computed from information in that slot. The list of possibilities is found by examining thedataanddataDerivedelements ofx[["?"]].Drop the
lonlimandlatlimparameters, marked for removal in 2014; useclongitude,clatitudeandspaninstead (seeplot,coastline-method()).
February 2016:
Drop the
fillparameter for land colour; usecolCoastlineinstead.Add the
borderCoastlineargument, to control the colour of coastlines and international boundaries.
See also
The documentation for ctd explains the structure of CTD objects, and also outlines the other functions dealing with them.
Other functions that plot oce data:
plot,adp-method,
plot,adv-method,
plot,amsr-method,
plot,argo-method,
plot,bremen-method,
plot,cm-method,
plot,coastline-method,
plot,gps-method,
plot,ladp-method,
plot,landsat-method,
plot,lisst-method,
plot,lobo-method,
plot,met-method,
plot,odf-method,
plot,rsk-method,
plot,satellite-method,
plot,sealevel-method,
plot,section-method,
plot,tidem-method,
plot,topo-method,
plot,windrose-method,
plot,xbt-method,
plotProfile(),
plotScan(),
plotTS()
Other things related to ctd data:
CTD_BCD2014666_008_1_DN.ODF.gz,
[[,ctd-method,
[[<-,ctd-method,
argo2ctd(),
as.ctd(),
cnvName2oceName(),
ctd,
ctd-class,
ctd.cnv.gz,
ctdDecimate(),
ctdFindProfiles(),
ctdFindProfilesRBR(),
ctdRaw,
ctdRepair(),
ctdTrim(),
ctd_aml_type1.csv.gz,
ctd_aml_type3.csv.gz,
d200321-001.ctd.gz,
d201211_0011.cnv.gz,
handleFlags,ctd-method,
initialize,ctd-method,
initializeFlagScheme,ctd-method,
oceNames2whpNames(),
oceUnits2whpUnits(),
plotProfile(),
plotScan(),
plotTS(),
read.ctd(),
read.ctd.aml(),
read.ctd.itp(),
read.ctd.odf(),
read.ctd.odv(),
read.ctd.saiv(),
read.ctd.sbe(),
read.ctd.ssda(),
read.ctd.woce(),
read.ctd.woce.other(),
setFlags,ctd-method,
subset,ctd-method,
summary,ctd-method,
woceNames2oceNames(),
woceUnit2oceUnit(),
write.ctd()
Examples
# 1. simple plot
library(oce)
data(ctd)
plot(ctd)
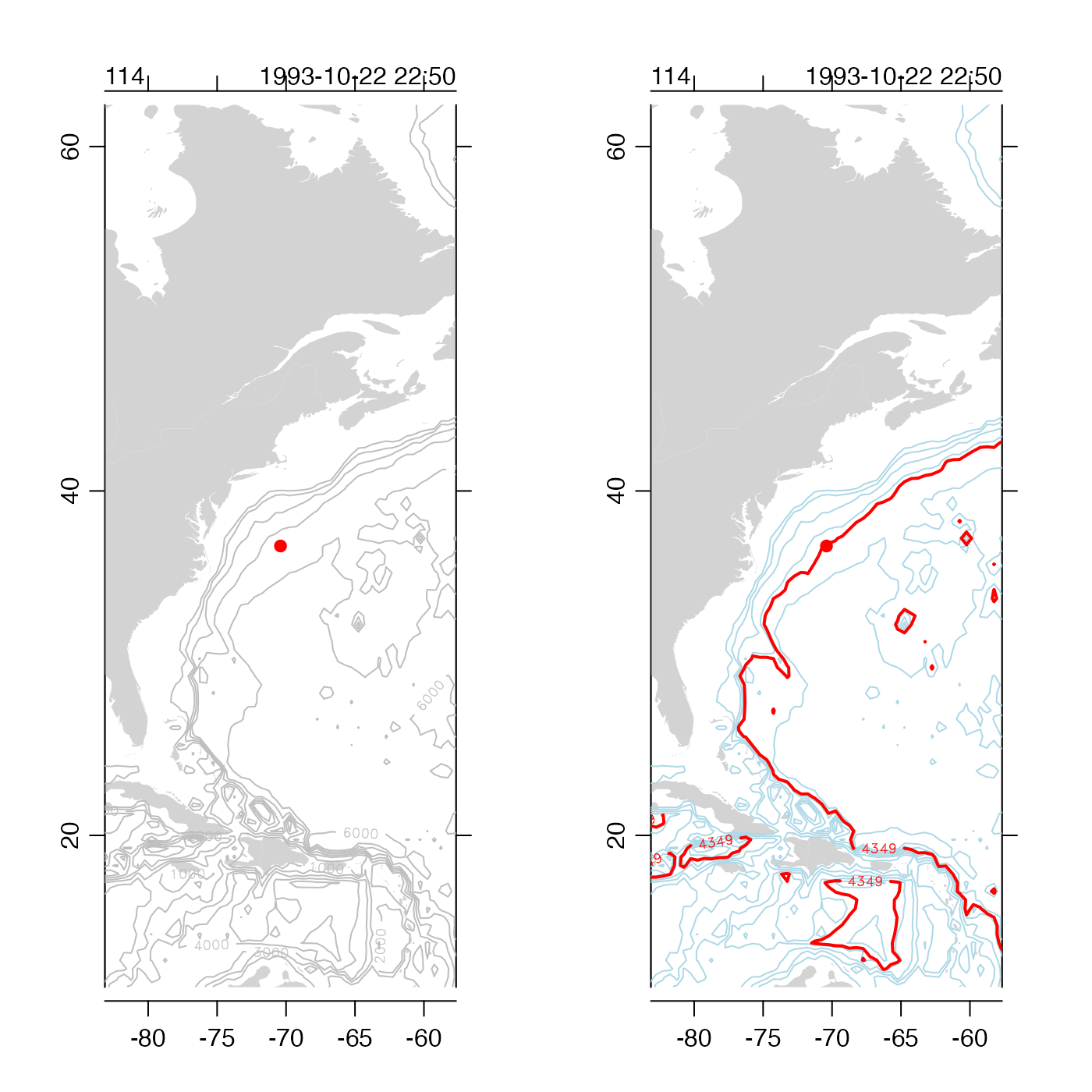
 # 2. how to customize depth contours
par(mfrow = c(1, 2))
data(section)
stn <- section[["station", 105]]
plot(stn, which = "map", drawIsobaths = TRUE)
plot(stn, which = "map")
data(topoWorld)
tlon <- topoWorld[["longitude"]]
tlat <- topoWorld[["latitude"]]
tdep <- -topoWorld[["z"]]
contour(tlon, tlat, tdep,
drawlabels = FALSE,
levels = seq(1000, 6000, 1000), col = "lightblue", add = TRUE
)
contour(tlon, tlat, tdep,
vfont = c("sans serif", "bold"),
levels = stn[["waterDepth"]], col = "red", lwd = 2, add = TRUE
)
# 2. how to customize depth contours
par(mfrow = c(1, 2))
data(section)
stn <- section[["station", 105]]
plot(stn, which = "map", drawIsobaths = TRUE)
plot(stn, which = "map")
data(topoWorld)
tlon <- topoWorld[["longitude"]]
tlat <- topoWorld[["latitude"]]
tdep <- -topoWorld[["z"]]
contour(tlon, tlat, tdep,
drawlabels = FALSE,
levels = seq(1000, 6000, 1000), col = "lightblue", add = TRUE
)
contour(tlon, tlat, tdep,
vfont = c("sans serif", "bold"),
levels = stn[["waterDepth"]], col = "red", lwd = 2, add = TRUE
)