Filter a time-series, possibly recursively
Arguments
- x
a vector of numeric values, to be filtered as a time series.
- a
a vector of numeric values, giving the \(a\) coefficients (see “Details”).
- b
a vector of numeric values, giving the \(b\) coefficients (see “Details”).
- zero.phase
boolean, set to
TRUEto run the filter forwards, and then backwards, thus removing any phase shifts associated with the filter.
Details
The filter is defined as e.g. \(y[i]=b[1]*x[i] + \)\( b[2]*x[i-1] + b[3]*x[i-2] + ... - a[2]*y[i-1] - a[3]*y[i-2] -
\)\( a[4]*y[i-3] - ...\), where some of the illustrated terms will be omitted if the lengths of
a and b are too small, and terms are dropped at the start of
the time series where the index on x would be less than 1.
By contrast with the filter() function of R, oce.filter
lacks the option to do a circular filter. As a consequence,
oceFilter introduces a phase lag. One way to remove this lag is to
run the filter forwards and then backwards, as in the “Examples”.
However, the result is still problematic, in the sense that applying it in
the reverse order would yield a different result. (Matlab's filtfilt
shares this problem.)
Note
The first value in the a vector is ignored, and if
length(a) equals 1, a non-recursive filter results.
Examples
library(oce)
par(mar = c(4, 4, 1, 1))
b <- rep(1, 5) / 5
a <- 1
x <- seq(0, 10)
y <- ifelse(x == 5, 1, 0)
f1 <- oceFilter(y, a, b)
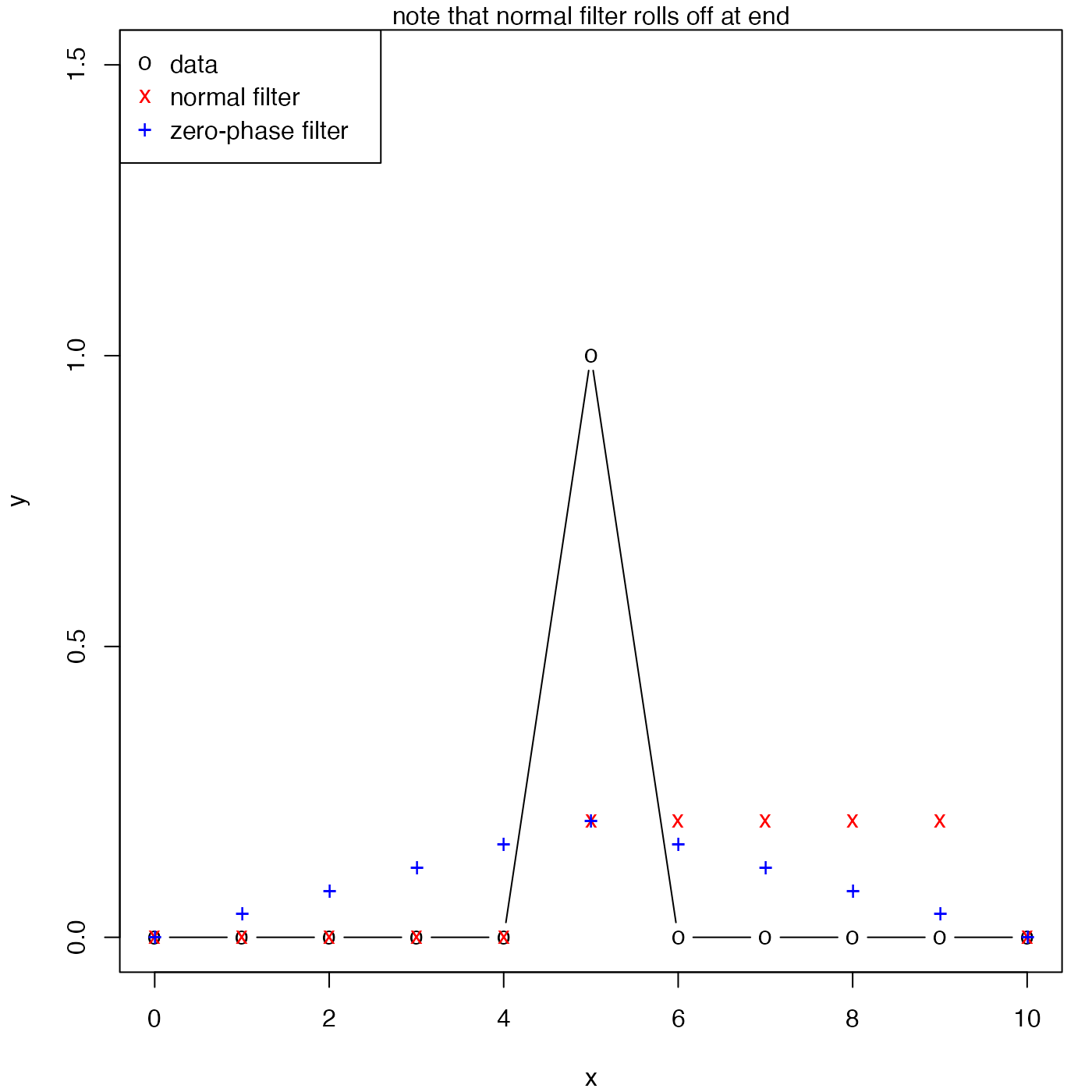
plot(x, y, ylim = c(-0, 1.5), pch = "o", type = "b")
points(x, f1, pch = "x", col = "red")
# remove the phase lag
f2 <- oceFilter(y, a, b, TRUE)
points(x, f2, pch = "+", col = "blue")
legend("topleft",
col = c("black", "red", "blue"), pch = c("o", "x", "+"),
legend = c("data", "normal filter", "zero-phase filter")
)
mtext("note that normal filter rolls off at end")